ACHAIKI IATRIKI | 2021; 40(3):133–136
Editorial
Evanthia Tourkochristou1,2, Athanasia Mouzaki1,3
1Division of Hematology, Department of Internal Medicine, Medical School, University of Patras, Patras, Greece
2Division of Gastroenterology, Department of Internal Medicine, Medical School, University of Patras, Patras, Greece
3Laboratory of Molecular Diagnosis of Infectious Agents, Medical School, University of Patras, Patras, Greece
Received: 30 Mar 2021; Accepted: 12 May 2021
Corresponding author: Athanasia Mouzaki, Laboratory of Molecular Diagnosis of Infectious Agents, Medical School, University of Patras, Patras, GR-26500, Greece, Tel.: +30 2610-969123, ORCID: 0000-0001-5548-7002, E-mail: mouzaki@upatras.gr
Key words: SARS-CoV-2; gastrointestinal tract; liver; ACE2; cytokines; inflammation
Introduction
The COVID-19 pandemic has posed major public health challenges worldwide. Severe acute respiratory syndrome coronavirus-2 (SARS-CoV-2) is transmitted primarily through the respiratory tract and causes typical flu-like symptoms of mild to moderate severity. However, evidence is accumulating that SARS-CoV-2 can also affect the gastrointestinal (GI) tract. An analysis of clinical data from 4,434 COVID-19 patients showed that the pooled prevalence of GI manifestations was 11.51% of infected patients. The most common symptom was diarrhea (7.78%), followed by nausea/vomiting (3.57%), loss of appetite (2.39%), and abdominal pain/discomfort (0.78%) [1]. In addition, autopsy of COVID-19 patients and imaging studies revealed microscopic and macroscopic changes and abnormalities of gastrointestinal tissues, including segmental dilatation and stenosis of the small intestine combined with mucosal detachment and necrosis, and colitis/enteritis characterized by inflammatory infiltrates and interstitial edema [2]. Mild and transient liver injury characterized by abnormal levels of alanine aminotransferase, aspartate aminotransferase, alkaline phosphatase, and mildly elevated serum bilirubin was noted in COVID-19 patients, with a greater extent of liver injury observed in patients with severe disease. Pathological liver tissue findings observed include macrovesicular and microvesicular steatosis, histiocytic hyperplasia, mild hepatic lobular and portal tract inflammatory infiltrates, increased platelet-fibrin microthrombi in the hepatic sinusoids, central or portal vein, rare megakaryocytes in the sinusoids, and hepatic necrosis of the ischemic type [3]. The prolonged presence (mean 11 days) of SARS-CoV-2 mRNA in fecal samples in more than half of COVID-19 patients after clearance of respiratory samples underscored the need for further investigation of the GI tract as another potent route of viral transmission [4].
SARS-CoV-2 transmission in the GI tract
A proposed mechanism of SARS-CoV-2 transmission in the digestive tract is shown in Figure 1. SARS-Cov-2 invades host cells using the spike (S) glycoprotein, which consists of an S1 receptor-binding domain (RBD) and an S2 domain. Fusion of viral and host cell membranes depends on activation of the S protein mediated by the proteases furin and transmembrane serine 2 (TMPRSS2), which cleave the S protein at two sites (S1/S2 and S2) and stimulate the release of the fusion peptide from the virus. Furin-mediated cleavage at the S1/S2 site can cause conformational changes in the viral protein, making the RBD and S2 domains accessible. The S protein binds to angiotensin converting enzyme 2 (ACE2) via the S1 receptor binding domain. Removal of the S1 domain from the viral surface allows the S2 domain to fuse with the host cell membrane, allowing viral entry [5].
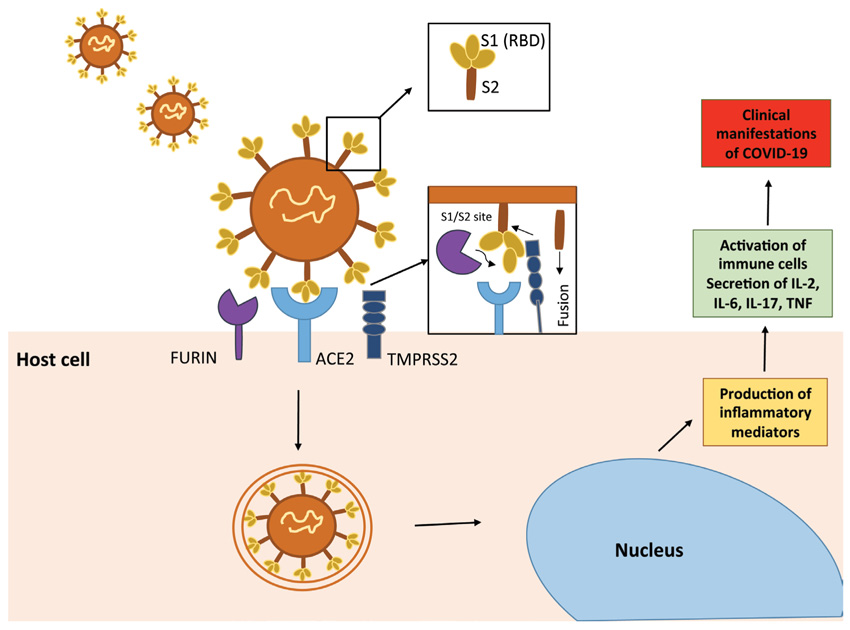
Figure 1. A proposed mechanism of SARS-CoV-2 transmission to the digestive tract. SARS-CoV-2 invades host cells using the spike (S) glycoprotein, which consists of an S1 receptor-binding domain (RBD) and an S2 domain. Fusion of viral and host cell membranes depends on activation of the S protein mediated by the proteases furin and transmembrane serine 2 (TMPRSS2), which cleave the S protein at two sites (S1/S2 and S2) and induce release of the fusion peptide (S2) from the virus. The S protein binds to a metallopeptidase, angiotensin converting enzyme-2 (ACE2), via the S1 receptor binding domain. When the S1 domain is removed from the viral surface, the S2 domain can fuse with the host cell membrane, allowing entry of the virus. Upon infection of ACE2-expressing cells, SARS-CoV-2 is thought to stimulate the production of inflammatory mediators, which in turn activate immune cells. The release of inflammatory cytokines (IL-2, IL-6, IL-17, TNF) by activated immune cells contributes to the clinical manifestations of COVID-19.
Upon infection of ACE2-expressing cells, SARS-CoV-2 is thought to stimulate the production of inflammatory mediators, which in turn activate immune cells. The release of inflammatory cytokines (IL-2, IL-6, IL-17, TNF) by activated immune cells contributes to the clinical manifestations of COVID-19 [6]. ACE2 is highly expressed in upper esophageal epithelial cells and small intestinal enterocytes, and higher expression of ACE2 was found in absorptive enterocytes of the ileum and colon compared to lung tissue. Single-cell transcriptome studies have shown that ACE2 and TMPRSS2 are highly co-expressed in upper esophageal cells, upper epithelial and glandular cells, and enterocytes of the ileum and colon, allowing viral invasion of the digestive tract [7].
COVID-19 pathophysiology of the digestive system
Effect of SARS-CoV-2 on the GI tract
The relatively high expression of viral receptors in cells of the GI tract makes the digestive system susceptible to SARS CoV-2 infection. The mechanism underlying the manifestation of GI symptoms in COVID-19 is not clear and several theories have been proposed. Most human coronaviruses have evolved through the presence of structural similarities and interspecies immunological cross-reactivity between animal and human coronaviruses, some of which can inherently cause gastroenteritis and maintain their enteric infectivity through interspecies recombination events, suggesting the possible gastrointestinal activity of SARS-CoV-2 [8], which can exert its effect on enterocytes by binding to ACE2. SARS-CoV-2 is thought to contribute to intestinal inflammation by interfering with the ACE2-dependent absorption of tryptophan, a major component of antimicrobial peptides. Decreased absorption of tryptophan due to ACE2 occupation by SARS-CoV-2 may lead to decreased absorption of antimicrobial peptides, promoting disruption of gut microbiota homeostasis and triggering of inflammation [9]. Alteration of the gut microbiota has been reported in COVID-19 patients, with dysbiosis of the gut microbiota persisting after resolution of the disease. This could reinforce the presence of intestinal symptomatology, considering the potent role of the gut microbiome in modulating immune responses and an association between gut microbiota composition and levels of cytokines and inflammatory markers found in patients with COVID-19 [10]. Α possible direct viral attack on enterocytes may lead to cellular dysfunction and increased permeability, which could be responsible for malabsorption and diarrhea. Symptoms of nausea and vomiting could be the result of a similar mechanism, considering the ability of SARS-CoV-2 to invade the upper GI glandular epithelium [7]. Damage to the GI tract could be part of the observed acute systemic inflammatory response and multi-organ failure in COVID-19, with dysregulation of cytokine levels and abnormal immune responses exacerbating the severity of the disease. High serum levels of pro-inflammatory cytokines and chemoattractant molecules, including IFN-γ, IL-6, IFN-γ-inducible protein-10 (IP-10), monocyte chemoattractant protein-1 (MCP-1), human granulocyte-macrophage colony-stimulating factor (GMCSF), are found in COVID-19 patients. In addition, activated T cells were found in the peripheral blood of a COVID-19 patient, showing high cytotoxicity with increased levels of cytotoxic granules granulysin and perforin, indicating a possible association of pathogenic T cells with the manifestation of systemic inflammation [11,12].
The distribution of viral RNA and nucleocapsid protein in the gastric, duodenal, and rectal epithelium of COVID-19 patients was also observed [13], suggesting that the virus can replicate and persist in the GI tract. A higher prevalence of GI symptoms, hypoxia-induced necrosis, and cellular injury leading to enterocyte dysfunction has been associated with various drugs administered to COVID-19 patients, including antiviral agents, antibiotics, and immunomodulators [2]. It is not yet clear whether the intestinal lesions in COVID-19 occur after systemic inflammation as a result of a secondary reaction or are caused by a primary intestinal infection or a combination of the above factors, and further analysis of intestinal samples from biopsy and autopsy should be performed. It should be noted that fecal calprotectin, which is a biomarker of intestinal inflammation in inflammatory bowel disease and infectious colitis, was found elevated in hospitalized COVID-19 patients who had resolved or persistent diarrhea compared with patients without diarrhea. This is a finding that should be considered when treating patients with preexisting gastrointestinal disease who have already developed intestinal inflammation [14].
Effect of SARS-CoV-2 on the liver
Liver injury could be due to either immune response-related injury from direct viral infection of hepatocytes or to strong ACE2-dependent viral invasion into cholangiocytes, which could lead to dysregulation of liver function or drug-induced hepatotoxicity [15]. ACE2 is highly expressed in cholangiocytes and hepatocytes and physiologically contributes to the reduction of liver injury caused by the renin-angiotensin system by degrading angiotensin Ang II to Ang 1-7 [16]. The liver also contains a large number of immune cells, and the antibody-dependent enhancement of infection (ADE) and systemic inflammatory response/cytokine storm might be related to the immune capacity of the liver and enhance the deleterious effects of abnormal immune responses and inflammation in COVID-19. ADE is induced by antibodies produced against SARS-CoV-2 spike protein and promotes SARS-CoV-2 entry into immune cells and immune-mediated tissue damage [17]. Patients with severe COVID-19 show swelling and steatosis of hepatocytes, hyperplasia of Kupffer cells, mild proliferation of hepatic sinusoidal cells, and lymphocyte infiltration, as well as increased levels of the immune activation markers IL-2 receptor and IL-6, which have been correlated with disease severity [18]. Hypoxia and hypotension associated with acute respiratory syndrome could also contribute to liver injury or even liver failure in critically ill patients, as hypoxia is associated with an oxidative stress response and increased release of reactive oxygen species, which can trigger the production of various pro-inflammatory factors that cause liver injury [19].
Pancreatic injury characterized by amylase or lipase abnormalities is another GI manifestation of COVID-19, considering that ACE2 is highly expressed in pancreatic islet cells. SARS-CoV-2 could likely cause islet cell damage, leading to an increased risk of diabetes. Similar to the mechanisms of liver injury, the direct cytopathic effect of SARS-CoV-2 or the acute systemic inflammatory response and subsequent immune cell response and cytokine storm as well as COVID-19 medication could be responsible for pancreatic tissue damage and enzyme abnormalities [20].
Conclusions
COVID-19 disease presents with a wide spectrum of clinical manifestations in the digestive system, combined with abnormal laboratory and imaging findings. The ability of SARS-CoV-2 to invade host cells of the digestive system and the persistence of the virus in fecal samples from COVID-19 patients even after respiratory symptoms have resolved should not be overlooked, as this indicates possible oral-fecal transmission of the virus. The pathophysiological mechanisms of SARS-CoV-2 infection that underlie gastrointestinal symptoms have not yet been elucidated, highlighting the need for further studies, including the analysis of patients’ clinical samples (autopsies, biopsies) and imaging findings during different phases of the disease. In addition, experimental studies will lead to a better understanding of the cellular and molecular biology of the interaction of the virus with host cells and its effects on the immune system. From a clinical point of view, attention should also be paid to the monitoring of patients with pre-existing pathology in the digestive system to prevent exacerbation of the disease and severe complications.
Conflict of interest disclosure
None to declare.
Declaration of funding sources
ET is a recipient of KARATHEODORIS grant #80672 from the University of Patras.
References
1. Merola E, Armelao F, de Pretis G. Prevalence of gastrointestinal symptoms in coronavirus disease 2019: a meta-analysis. Acta Gastroenterol Belg. 2020; 83(4):603-15.
2. Tian Y, Rong L, Nian W, He Y. Review article: gastrointestinal features in COVID-19 and the possibility of faecal transmission. Aliment Pharmacol Ther. 2020; 51(9):843-51.
3. Zhao CL, Rapkiewicz A, Maghsoodi-Deerwester M, Gupta M, Cao W, Palaia T, et al. Pathological findings in the postmortem liver of patients with coronavirus disease 2019 (COVID-19). Hum Pathol. 2021; 109:59-68.
4. Wu Y, Guo C, Tang L, Hong Z, Zhou J, Dong X, et al. Prolonged presence of SARS-CoV-2 viral RNA in faecal samples: Lancet Gastroenterol Hepatol. 2020; 5(5): 434-5.
5. Shang J, Wan Y, Luo C, Ye G, Geng Q, Auerbach A, et al. Cell entry mechanisms of SARS-CoV-2. Proc Natl Acad Sci U S A. 2020; 117(21):11727-34.
6. Lin L, Lu L, Cao W, Li T. Hypothesis for potential pathogenesis of SARS-CoV-2 infection-a review of immune changes in patients with viral pneumonia. Emerg Microbes Infect. 2020; 9(1):727-32.
7. Zhang H, Kang Z, Gong H, Xu D, Wang J, Li Z, et al. The digestive system is a potential route of 2019-nCov infection: a bioinformatics analysis based on single-cell transcriptomes. BioRxiv. 2020.
8. Cimolai N. Features of enteric disease from human coronaviruses: Implications for COVID-19. J Med Virol. 2020; 92(10):1834-44.
9. Hashimoto T, Perlot T, Rehman A, Trichereau J, Ishiguro H, Paolino M, et al. ACE2 links amino acid malnutrition to microbial ecology and intestinal inflammation. Nature. 2012; 487(7408):477-81.
10. Yeoh YK, Zuo T, Lui GC, Zhang F, Liu Q, Li AY, et al. Gut microbiota composition reflects disease severity and dysfunctional immune responses in patients with COVID-19. Gut. 2021; 70(4):698-706.
11. Huang C, Wang Y, Li X, Ren L, Zhao J, Hu Y, et al. Clinical features of patients infected with 2019 novel coronavirus in Wuhan, China. Lancet. 2020; 395(10223):497-506.
12. Xu Z, Shi L, Wang Y, Zhang J, Huang L, Zhang C, et al. Pathological findings of COVID-19 associated with acute respiratory distress syndrome: Lancet Respir Med. 2020; 8(4):420-2.
13. Xiao F, Tang M, Zheng X, Liu Y, Li X, Shan H. Evidence for Gastrointestinal Infection of SARS-CoV-2. Gastroenterology. 2020; 158(6):1831-3.
14. Effenberger M, Grabherr F, Mayr L, Schwaerzler J, Nairz M et al. Faecal calprotectin indicates intestinal inflammation in COVID-19. Gut. 2020; 69(8):1543-4.
15. Xu L, Liu J, Lu M, Yang D, Zheng X. Liver injury during highly pathogenic human coronavirus infections. Liver Int. 2020; 40(5):998-1004.
16. Paizis G, Tikellis C, Cooper ME, Schembri JM, Lew RA, Smith AI, et al. Chronic liver injury in rats and humans upregulates the novel enzyme angiotensin converting enzyme 2. Gut. 2005; 54(12):1790-6.
17. Wang SF, Tseng SP, Yen CH, Yang JY, Tsao CH, Shen CW, et al. Antibody-dependent SARS coronavirus infection is mediated by antibodies against spike proteins. Biochem Biophys Res Commun. 2014; 451(2):208-14.
18. Zhu N, Zhang D, Wang W, Li X, Yang B, Song J, et al. A Novel Coronavirus from Patients with Pneumonia in China, 2019. N Engl J Med. 2020; 382(8):727-33.
19. Chand N, Sanyal AJ. Sepsis-induced cholestasis. Hepatology. 2007; 45(1):230-41.
20. Wang F, Wang H, Fan J, Zhang Y, Zhao Q. Pancreatic Injury Patterns in Patients With Coronavirus Disease 19 Pneumonia. Gastroenterology. 2020; 159(1):367-70.